Porphyromonas gingivalis: Difference between revisions
imported>John J. Dennehy |
imported>Yelena Fuzaylova |
||
| Line 34: | Line 34: | ||
'''Carbohydrate Metabolism:''' | '''Carbohydrate Metabolism:''' | ||
Glycolysis / Gluconeogenesis | Glycolysis / Gluconeogenesis | ||
| Line 62: | Line 63: | ||
Oxidative phosphorylation | Oxidative phosphorylation | ||
ATP synthase | ATP synthase | ||
Methane metabolism | Methane metabolism | ||
Carbon fixation | Carbon fixation | ||
Reductive carboxylate cycle (CO2 fixation) | Reductive carboxylate cycle (CO2 fixation) | ||
Nitrogen metabolism | Nitrogen metabolism | ||
Sulfur metabolism | Sulfur metabolism | ||
| Line 72: | Line 79: | ||
Fatty acid biosynthesis | Fatty acid biosynthesis | ||
Fatty acid metabolism | Fatty acid metabolism | ||
Synthesis and degradation of ketone bodies | Synthesis and degradation of ketone bodies | ||
Sterol biosynthesis | Sterol biosynthesis | ||
Bile acid biosynthesis | Bile acid biosynthesis | ||
C21-Steroid hormone metabolism | C21-Steroid hormone metabolism | ||
Androgen and estrogen metabolism | Androgen and estrogen metabolism | ||
| Line 82: | Line 95: | ||
Purine metabolism | Purine metabolism | ||
Pyrimidine metabolism | Pyrimidine metabolism | ||
Nucleotide sugars metabolism | Nucleotide sugars metabolism | ||
| Line 88: | Line 103: | ||
Glutamate metabolism | Glutamate metabolism | ||
Alanine and aspartate metabolism | Alanine and aspartate metabolism | ||
Glycine, serine and threonine metabolism | Glycine, serine and threonine metabolism | ||
Methionine metabolism | Methionine metabolism | ||
Cysteine metabolism | Cysteine metabolism | ||
Valine, leucine and isoleucine degradation | Valine, leucine and isoleucine degradation | ||
Valine, leucine and isoleucine biosynthesis | Valine, leucine and isoleucine biosynthesis | ||
Lysine biosynthesis | Lysine biosynthesis | ||
Lysine degradation | Lysine degradation | ||
Arginine and proline metabolism | Arginine and proline metabolism | ||
Histidine metabolism | Histidine metabolism | ||
Tyrosine metabolism | Tyrosine metabolism | ||
Phenylalanine metabolism | Phenylalanine metabolism | ||
Tryptophan metabolism | Tryptophan metabolism | ||
Phenylalanine, tyrosine and tryptophan biosynthesis | Phenylalanine, tyrosine and tryptophan biosynthesis | ||
Urea cycle and metabolism of amino groups | Urea cycle and metabolism of amino groups | ||
| Line 107: | Line 137: | ||
beta-Alanine metabolism | beta-Alanine metabolism | ||
Taurine and hypotaurine metabolism | Taurine and hypotaurine metabolism | ||
Aminophosphonate metabolism | Aminophosphonate metabolism | ||
Selenoamino acid metabolism | Selenoamino acid metabolism | ||
Cyanoamino acid metabolism | Cyanoamino acid metabolism | ||
D-Glutamine and D-glutamate metabolism | D-Glutamine and D-glutamate metabolism | ||
D-Arginine and D-ornithine metabolism | D-Arginine and D-ornithine metabolism | ||
D-Alanine metabolism | D-Alanine metabolism | ||
Glutathione metabolism | Glutathione metabolism | ||
'''Metabolism of Complex Carbohydrates:''' | '''Metabolism of Complex Carbohydrates:''' | ||
Starch and sucrose metabolism | Starch and sucrose metabolism | ||
Biosynthesis and degradation of glycoprotein | Biosynthesis and degradation of glycoprotein | ||
Aminosugars metabolism | Aminosugars metabolism | ||
Lipopolysaccharide biosynthesis | Lipopolysaccharide biosynthesis | ||
Peptideglycan biosynthesis | Peptideglycan biosynthesis | ||
Glycosaminoglycan degradation | Glycosaminoglycan degradation | ||
'''Metabolism of Complex Lipids:''' | '''Metabolism of Complex Lipids:''' | ||
Glycerolipid metabolism | Glycerolipid metabolism | ||
Inositol phosphate metabolism | Inositol phosphate metabolism | ||
Sphingophospholipid biosynthesis | Sphingophospholipid biosynthesis | ||
Phospholipid degradation | Phospholipid degradation | ||
Sphingoglycolipid metabolism | Sphingoglycolipid metabolism | ||
Prostaglandin and leukotriene metabolism | Prostaglandin and leukotriene metabolism | ||
| Line 135: | Line 185: | ||
Thiamine metabolism | Thiamine metabolism | ||
Riboflavin metabolism | Riboflavin metabolism | ||
Vitamin B6 metabolism | Vitamin B6 metabolism | ||
Nicotinate and nicotinamide metabolism | Nicotinate and nicotinamide metabolism | ||
Pantothenate and CoA biosynthesis | Pantothenate and CoA biosynthesis | ||
Biotin metabolism | Biotin metabolism | ||
Folate biosynthesis | Folate biosynthesis | ||
One carbon pool by folate | One carbon pool by folate | ||
Retinol metabolism | Retinol metabolism | ||
Porphyrin and chlorophyll metabolism | Porphyrin and chlorophyll metabolism | ||
Ubiquinone biosynthesis | Ubiquinone biosynthesis | ||
Revision as of 19:36, 1 April 2008
Articles that lack this notice, including many Eduzendium ones, welcome your collaboration! |
Classification
Higher order taxa
Domain:Bacteria
Phylum:Bacteroidetes
Class:Bacteroides
Genus:Porphyromonas gingivalis
Species
Genus species:Porphyromonas gingivalis
Description and significance
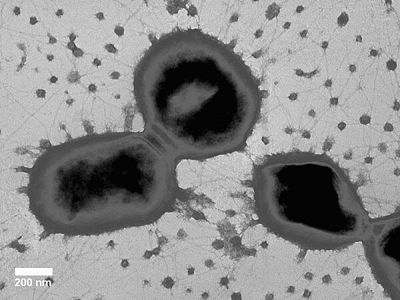
Porphyromonas gingivalis is an anaerobic, gram negative bacterium that can be found within the mouth of an individual. It is observed to be non-motile and rod-shaped. This bacterium is the principal source of periodontal disease. It has been found that in addition to causing human infections, this bacterium also causes much of the antibiotic resistance problems found today.[1] The way which it operates is very unique, since it is a gram negative bacteria, it can attach to the subgingival coating of the tooth, and it will substitute the gram positive bacteria that is originally there with its own thus causing an inflammation which will disengage the gums from the teeth. When colonized on blood agar it forms black spots. It is important to sequence the genome of this organism because it is found in many locations within the body not only in the oral cavity also in the gastrointestinal tract, respiratory tract and in the colon, this has many consequences. It has been ascertain that periodontal disease is associated with cardiovascular diseases. This bacterium is associated with Treptonema denticola and Bacteroides forsythus [2]
Genome structure
The genome of p. gingivalis is circular and its origin of replication is located at oriC which is juxtaposed by the genes dnaA and PG1949. The guanine and cytosine nucleotides make up approximately 49%. At the present, 2,015 genes have been acknowledged with a total of 23,43,479 nucleotides. It has been found that the genome of p. gingivalis is similar to that of Bacteroides thetaiotaomicron, and B.fragilis within the Cytophaga-Flavobacteria-Bacteroides genome. In addition, it closely resembles the genomes of Chlorobium tepidum, and thus this demonstrates the notion that the phyla of Chlorobia and Cytophaga-Flavobacteria-Bacteroides are associated with one another. It has been discovered using genome analysis that this bacterium can metabolize a range of amino acids which will form different metabolic end products that are lethal to the host which is usually human, as well as harming the host’s gingival tissues, thus causing the expansion of periodontal disease. [3] It is found that the size of the genome of P. gingivalis is 2.34 mega bases. In addition, the sequencing of the entire genome is important to be able to determine if there are more medical effects of the bacterium in addition to periodontal disease. Researchers want to be able to establish mechanisms for the virulence as well as vaccines. The goals of this project include, the DNA sequence for the genome; analyzing and interpreting the sequence for the W83 strain; integrate the information that was obtained by the interpreted sequences and compare it with the experimental data from the P. gingivalis research community; finally the purpose of the genome sequences is to be able to make clones, reagents, and information to the research community available. (http://www.pgingivalis.org/aims.htm)
Cell structure and metabolism
The P.gingivalis is a gram-negative, non-motile, rod-shaped, anaerobic organism. To function, it undergoes a mechanism in which it binds to the subgingival layer of the mouth using fimbriae. These fimbriae not only aid in the role of adhesion but are also found to be pathogenic to the immune system. [4] P. gingivalis takes part in Iron Transport, the way it does this is by using a hemin as a device to help it transport iron. When this builds up it results in the black pigmentation that is detected. To bring the hemin iron complex into the cell it uses an ABC Transporter. In regards to the metabolism P. gingivalis, it can undergo many different types such as:
Carbohydrate Metabolism:
Glycolysis / Gluconeogenesis
Citrate cycle (TCA cycle)
Pentose phosphate cycle
Pentose and glucuronate interconversions
Fructose and mannose metabolism
Galactose metabolism
Ascorbate and aldarate metabolism
Pyruvate metabolism
Glyoxylate and dicarboxylate metabolism
Propanoate metabolism
Butanoate metabolism
C5-Branched dibasic acid metabolism
Energy Metabolism:
Oxidative phosphorylation
ATP synthase
Methane metabolism
Carbon fixation
Reductive carboxylate cycle (CO2 fixation)
Nitrogen metabolism
Sulfur metabolism
Lipid Metabolism:
Fatty acid biosynthesis
Fatty acid metabolism
Synthesis and degradation of ketone bodies
Sterol biosynthesis
Bile acid biosynthesis
C21-Steroid hormone metabolism
Androgen and estrogen metabolism
Nucleotide Metabolism:
Purine metabolism
Pyrimidine metabolism
Nucleotide sugars metabolism
Amino Acid Metabolism:
Glutamate metabolism
Alanine and aspartate metabolism
Glycine, serine and threonine metabolism
Methionine metabolism
Cysteine metabolism
Valine, leucine and isoleucine degradation
Valine, leucine and isoleucine biosynthesis
Lysine biosynthesis
Lysine degradation
Arginine and proline metabolism
Histidine metabolism
Tyrosine metabolism
Phenylalanine metabolism
Tryptophan metabolism
Phenylalanine, tyrosine and tryptophan biosynthesis
Urea cycle and metabolism of amino groups
Metabolism of Other Amino Acids:
beta-Alanine metabolism
Taurine and hypotaurine metabolism
Aminophosphonate metabolism
Selenoamino acid metabolism
Cyanoamino acid metabolism
D-Glutamine and D-glutamate metabolism
D-Arginine and D-ornithine metabolism
D-Alanine metabolism
Glutathione metabolism
Metabolism of Complex Carbohydrates:
Starch and sucrose metabolism
Biosynthesis and degradation of glycoprotein
Aminosugars metabolism
Lipopolysaccharide biosynthesis
Peptideglycan biosynthesis
Glycosaminoglycan degradation
Metabolism of Complex Lipids:
Glycerolipid metabolism
Inositol phosphate metabolism
Sphingophospholipid biosynthesis
Phospholipid degradation
Sphingoglycolipid metabolism
Prostaglandin and leukotriene metabolism
Metabolism of Cofactors and Vitamins:
Thiamine metabolism
Riboflavin metabolism
Vitamin B6 metabolism
Nicotinate and nicotinamide metabolism
Pantothenate and CoA biosynthesis
Biotin metabolism
Folate biosynthesis
One carbon pool by folate
Retinol metabolism
Porphyrin and chlorophyll metabolism
Ubiquinone biosynthesis
Ecology
The organism P. gingivalis is usually found in the orifice of mammals usually humans. It is associated with Treptonema denticola and Bacteroides forsythus. There is no valid evidence that the relationship is symbiotic, but is still under researched theory. It contributes to the formation of gingivitis which is a periodontal disease.
Pathology
How does this organism cause disease? Human, animal, plant hosts? Virulence factors, as well as patient symptoms.
Application to Biotechnology
Does this organism produce any useful compounds or enzymes? What are they and how are they used?
Current Research
Enter summaries of the most recent research here--at least three required